Pairwise Sequence Alignment
Pairwise sequence alignment involves comparing two biological sequences - typically DNA, RNA, or protein sequences - to identify regions of similarity that may indicate functional, structural, or evolutionary relationships between the sequences. Highly similar sequences can be considered members of the same family.
As a bioinformatics student, understanding the principles, algorithms, and applications of pairwise sequence alignment is crucial.
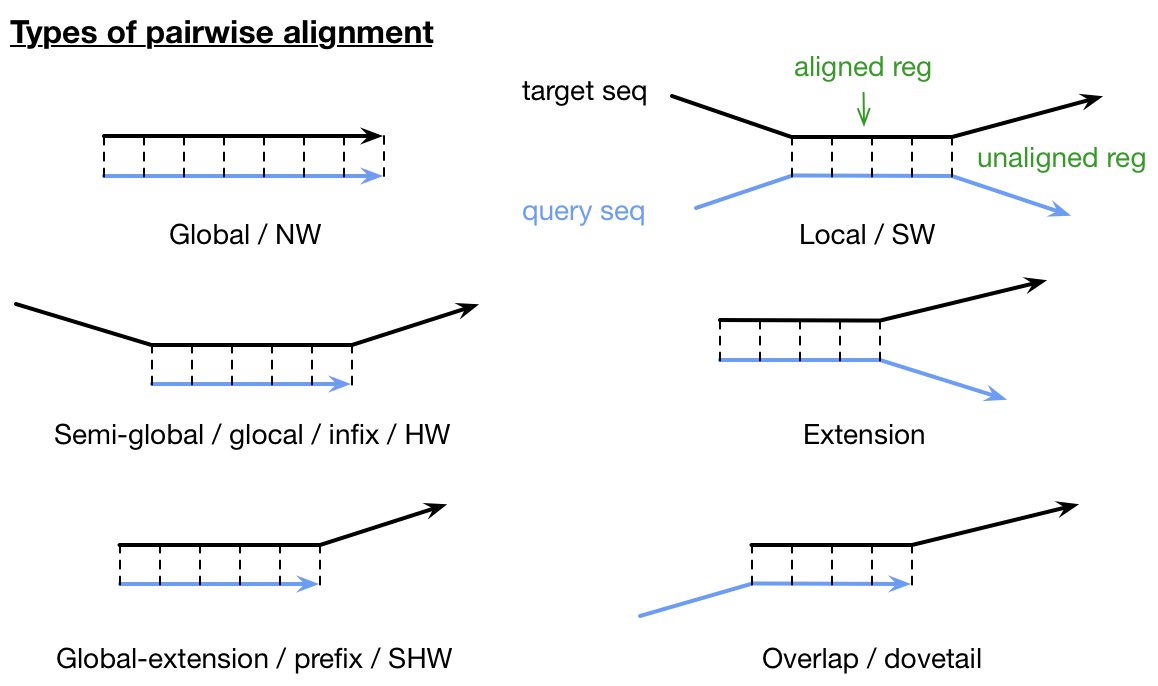
1. Types of Pairwise Alignment
Pairwise sequence alignment can be categorized into three main types:
-
Global Alignment: Attempts to align every residue in both sequences, spanning the entire length of each sequence. This is most useful when the sequences are similar and of roughly equal size. Global alignment provides complete sequence pairing and helps identify small variations and polymorphisms. Global alignment is not suitable for divergent sequences or sequences of significantly different lengths, as it may fail to identify highly similar local regions.
-
Local Alignment: Identifies regions of similarity within long sequences that are often widely divergent overall. This is more useful for dissimilar sequences that are suspected to contain regions of similarity or similar sequence motifs. It identifies subsequences with high similarity, regardless of their location within the sequences. This approach is suitable for finding evolutionarily conserved elements, gene pairs, repeats (including transposons), and other shared features. Local alignment is especially useful when sequences have similar parts but differ overall, as it ignores mismatches and focuses on aligned portions.
-
Semi-Global Alignment: Also known as free-end gaps alignment, this method does not penalize gaps at the beginning or end of either sequence. It’s useful when aligning sequences of different lengths or when searching for a shorter sequence within a longer one. It’s a modification of the global alignment algorithm where gaps at the beginning and end of either sequence are not penalized.
2. Dot-matrix method to align two sequences visually
Dot-matrix method is a technique for comparing two sequences by visually representing matching positions as dots on a grid. Diagonal lines of dots indicate regions of similarity between the sequences.Dot-matrix plots can identify insertions, deletions, repetitions, and inverted repeats. The method is simple to understand and implement, but can be computationally intensive for large sequences.

Dot plots have drawbacks: they can be noisy, lack clarity, and are difficult to interpret for large sequences. Dot plots can be improved by using a threshold value and window size to filter out random matches. Dot plots offer benefits: easy implementation, clear visualization, and identification of specific sequence features. One drawback is that dot matrix methods do not produce a traditional alignment or provide a statistical significance score.
3. Needleman-Wunsch Algorithm for Global Alignment
The Needleman-Wunsch algorithm is a dynamic programming approach for global sequence alignment. It guarantees finding the optimal alignment given a particular scoring system.
3.1. Key steps of Needleman-Wunsch Algorithm:
- Initialization: Create a matrix of size (m+1) x (n+1), where m and n are the lengths of the two sequences.
- Matrix Filling: Fill the matrix using the recurrence relation:
Where s(x_i, y_j) is the substitution score, and g is the gap penalty.F(i,j) = max(F(i-1,j-1) + s(x_i, y_j),F(i-1,j) + g,F(i,j-1) + g)
- Traceback: Start from the bottom-right cell and trace back to the top-left to reconstruct the optimal alignment.
Time Complexity: O(mn) Space Complexity: O(mn)

3.2. Python implementation (simplified) of Needleman-Wunsch Algorithm:
def needleman_wunsch(seq1, seq2, match_score=1, mismatch_score=-1, gap_penalty=-2): m, n = len(seq1), len(seq2) score_matrix = [[0 for j in range(n+1)] for i in range(m+1)]
# Initialize first row and column for i in range(m+1): score_matrix[i][0] = i * gap_penalty for j in range(n+1): score_matrix[0][j] = j * gap_penalty
# Fill the score matrix for i in range(1, m+1): for j in range(1, n+1): match = score_matrix[i-1][j-1] + (match_score if seq1[i-1] == seq2[j-1] else mismatch_score) delete = score_matrix[i-1][j] + gap_penalty insert = score_matrix[i][j-1] + gap_penalty score_matrix[i][j] = max(match, delete, insert)
# Traceback align1, align2 = "", "" i, j = m, n while i > 0 and j > 0: score_current = score_matrix[i][j] score_diagonal = score_matrix[i-1][j-1] score_up = score_matrix[i][j-1] score_left = score_matrix[i-1][j]
if score_current == score_diagonal + (match_score if seq1[i-1] == seq2[j-1] else mismatch_score): align1 = seq1[i-1] + align1 align2 = seq2[j-1] + align2 i -= 1 j -= 1 elif score_current == score_left + gap_penalty: align1 = seq1[i-1] + align1 align2 = '-' + align2 i -= 1 elif score_current == score_up + gap_penalty: align1 = '-' + align1 align2 = seq2[j-1] + align2 j -= 1
while i > 0: align1 = seq1[i-1] + align1 align2 = '-' + align2 i -= 1 while j > 0: align1 = '-' + align1 align2 = seq2[j-1] + align2 j -= 1
return (align1, align2, score_matrix[m][n])4. Smith-Waterman Algorithm for Local Alignment
The Smith-Waterman algorithm is a dynamic programming approach for local sequence alignment. It’s similar to Needleman-Wunsch but allows for the identification of similar regions within sequences.
4.1. Key differences from Needleman-Wunsch:
- The matrix is initialized with zeros.
- Negative scores in the matrix are replaced with zeros.
- Traceback starts from the highest score in the matrix and ends when a cell with zero is encountered.
Time Complexity: O(mn) Space Complexity: O(mn)

4.2. Python implementation (simplified) of Smith-Waterman Algorithm:
def smith_waterman(seq1, seq2, match_score=2, mismatch_score=-1, gap_penalty=-2): m, n = len(seq1), len(seq2) score_matrix = [[0 for j in range(n+1)] for i in range(m+1)] max_score = 0 max_pos = None
# Fill the score matrix for i in range(1, m+1): for j in range(1, n+1): match = score_matrix[i-1][j-1] + (match_score if seq1[i-1] == seq2[j-1] else mismatch_score) delete = score_matrix[i-1][j] + gap_penalty insert = score_matrix[i][j-1] + gap_penalty score_matrix[i][j] = max(0, match, delete, insert) if score_matrix[i][j] > max_score: max_score = score_matrix[i][j] max_pos = (i, j)
# Traceback align1, align2 = "", "" i, j = max_pos while score_matrix[i][j] != 0: score_current = score_matrix[i][j] score_diagonal = score_matrix[i-1][j-1] score_up = score_matrix[i][j-1] score_left = score_matrix[i-1][j]
if score_current == score_diagonal + (match_score if seq1[i-1] == seq2[j-1] else mismatch_score): align1 = seq1[i-1] + align1 align2 = seq2[j-1] + align2 i -= 1 j -= 1 elif score_current == score_left + gap_penalty: align1 = seq1[i-1] + align1 align2 = '-' + align2 i -= 1 elif score_current == score_up + gap_penalty: align1 = '-' + align1 align2 = seq2[j-1] + align2 j -= 1
return (align1, align2, max_score)5. Scoring Systems
The choice of scoring system is crucial in sequence alignment as it determines how matches, mismatches, and gaps are evaluated.
- Nucleic Acid Scoring: Simple; matches get high/positive values, mismatches get negative values. For example:
- Match: +1
- Mismatch: -1
- Gap: -2
- Amino Acid Scoring Complexity: More complex due to physicochemical properties.
- Amino acids with similar properties are more likely to be substituted.
- Substitutions with different properties can disrupt protein structure and function.
- Examples of Amino Acid Substitutions:
- Aromatic Amino Acids (Tryptophan, Tyrosine, Phenylalanine): Easily substitute for each other because of similar properties.
- Basic Amino Acids (Lysine, Arginine, Histidine): Also frequently substitute due to shared properties.
- Cysteine: Substitution is less common because it can form disulfide bonds, which are crucial for protein structure.
5.1 Scoring matrices for amino acids
There are two types of scoring matrices for amino acids: feature-dependent and empirical.
- Empirical matrices (PAM and BLOSUM) are preferred and based on substitution probabilities.
- A positive score indicates a higher frequency of substitution than expected by chance.
- A score of 0 indicates a frequency equal to chance.
- A negative score indicates a lower frequency of substitution than expected by chance.
- Position-Specific Scoring Matrices (PSSMs) is a feature-dependent custom scoring matrix that capture position-specific information about a family of sequences.
5.2 PAM (point accepted mutation)
PAM (point accepted mutation) was created by Margaret Dayhoff and colleagues in the 1970s to develop substitution matrices of amino acids from related proteins. PAM matrices are based on evolutionary distance between sequences, with higher numbers indicating greater distance.

The PAM-1 matrix is deduced by selecting closely related sequences with mutation frequencies corresponding to PAM-1. PAM matrices are symmetrical and derived from the global alignment method. The PAM score is calculated through a multi-step process, including normalizing expected residue frequencies and using a ""log odd score"" on a log scale.

5.2.2 Example
- Alignment Probability: The probability of two residues aligning randomly is represented by Pr(C,D), while the likelihood of them aligning due to evolutionary relationships (orthologs) is Po(C,D).
- Score Function: A score function measures the significance of an alignment using the ratio of evolutionary alignment probability to random alignment probability: ¼ log Po(C,D)/Pr(C,D).
- PAM Matrices: PAM (Point Accepted Mutation) matrices are used to calculate evolutionary distances between protein sequences, based on the concept of one mutation per 100 amino acids. PAM 1 represents one mutation per 100 amino acids. Higher PAM numbers indicate greater evolutionary divergence.
- PAM matrices are limited by the small variation in proteins used for their construction, their bias towards globular proteins, and their reliance on a single dataset.
5.3 BLOcks SUbstitution matrix (BLOSUM)
BLOSUM matrices were developed by Henikoff and Henikoff in 1992 to address limitations of PAM matrices. They are based on alignments of conserved protein sites from the ""BLOCKS"" database, a collection of around 2000 ungapped alignments from 500 protein families. BLOSUM derived from local multiple sequence alignments of distantly related proteins.
Unlike PAM matrices, BLOSUM matrices use actual percentage identity to construct the matrix. BLOSUM62 is a specific BLOSUM matrix representing alignments with no more than 62% identity.
BLOSUM numbers work in the opposite way to PAM numbers - lower BLOSUM numbers indicate more divergent sequences.

Comparison of BLOSUM and PAM: BLOSUM scores are comparable to PAM scores at various evolutionary distances:
- BLOSUM90 is similar to PAM100
- BLOSUM80 is similar to PAM120
- BLOSUM60 is similar to PAM160
- BLOSUM52 is similar to PAM200
- BLOSUM45 is similar to PAM250
Technical Note: The choice of scoring system can significantly impact the alignment results. For example, BLOSUM62 is often used as a default for protein sequence alignments, but BLOSUM80 might be more appropriate for closely related sequences, while BLOSUM45 could be better for more divergent sequences.
5.1. Gaps and gap penalties
Gaps represent insertions or deletions in sequences. They are crucial for aligning sequences that have diverged over time. Gap Penalties: is a system to control the introduction and length of gaps in alignments. The goal is to find biologically relevant alignments, not just maximize the number of gaps.
Types of Gap Penalties:
- Constant (Fixed): A fixed penalty for each gap, regardless of its length. This encourages fewer, larger gaps.
- Linear: The penalty increases proportionally to the length of the gap. This favors shorter gaps.
- Affine: The most common type, combining a gap opening penalty (for starting a gap) and a gap extension penalty (for extending a gap). This allows for greater flexibility in controlling gap size and frequency.
5.2. Factors Influencing Gap Penalty Choice:
- Relationship to Match/Mismatch Scores: Gap penalties must be high enough to ensure that only significant matches are favored.
- Goal of the Alignment: Finding closely related matches requires higher penalties, while finding distant matches requires lower penalties.
6. BLAST and FASTA
While Smith-Waterman guarantees finding the optimal local alignment, it’s computationally expensive for large-scale sequence comparisons. Heuristic algorithms like BLAST (Basic Local Alignment Search Tool) and FASTA provide faster alternatives.
BLAST algorithm overview:
- Seeding: Break the query sequence into short subsequences (words).
- Extension: Find exact matches of these words in the database and extend them.
- Evaluation: Score the extended alignments and return high-scoring ones.

Technical Note: BLAST uses various optimizations to speed up the search process, such as neighborhood words (for proteins) and two-hit algorithm (for nucleotides). Understanding these optimizations is crucial for interpreting BLAST results and fine-tuning searches.