1. Navigating NCBI Databases
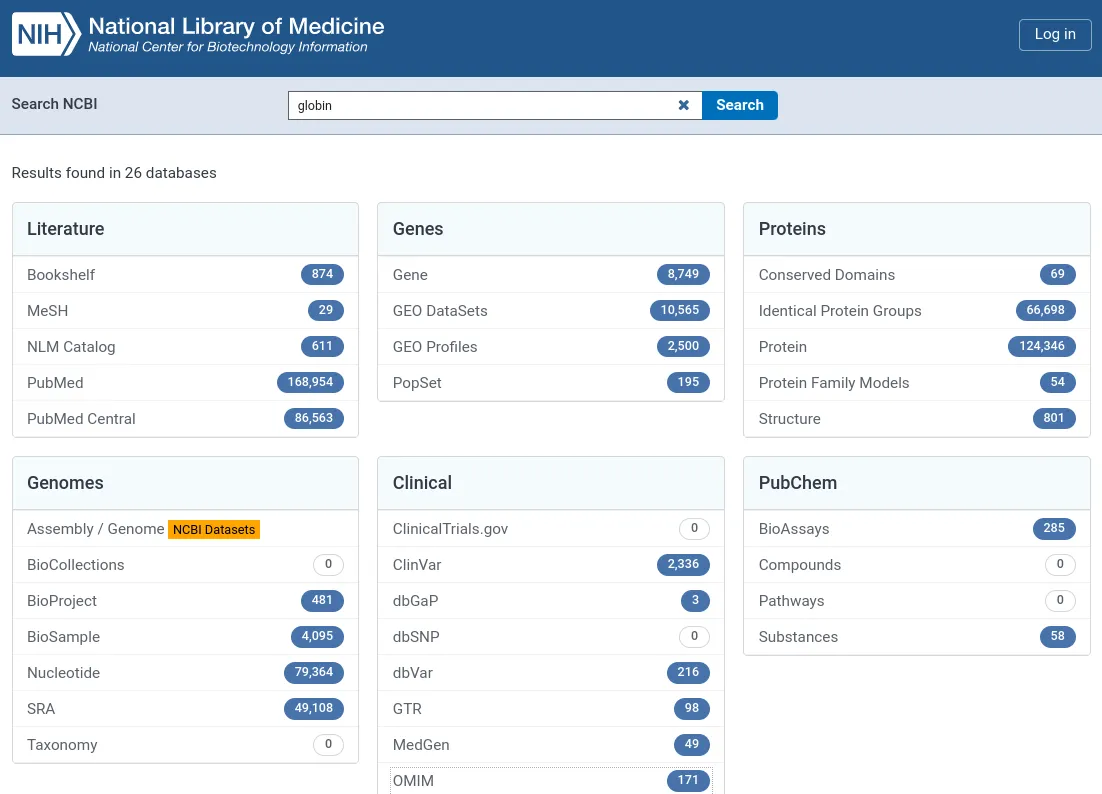
Visit https://www.ncbi.nlm.nih.gov/ and search for “globin”.
Section I. Literature Databases
1. PubMed (168,954 results)
- Purpose: Primary literature search for peer-reviewed articles
- Example: Search for recent studies on hemoglobin disorders
- Activity: Students find and summarize a recent paper on globin gene therapy
2. PubMed Central (86,563 results)
- Purpose: Free full-text archive of biomedical literature
- Example: Access and compare open-access articles on globin evolution
- Activity: Students identify key differences between PubMed and PubMed Central
3. Bookshelf (874 results)
- Purpose: Free access to books and documents in life sciences and healthcare
- Example: Find chapters on globin protein structure and function
- Activity: Students create a brief presentation on globin types using Bookshelf resources
4. MeSH (29 results)
- Purpose: Medical Subject Headings for indexing and searching biomedical literature
- Example: Explore MeSH terms related to globins (e.g., “Globins”, “Hemoglobins”, “Myoglobin”)
- Activity: Students create a MeSH tree for globin-related terms
5. NLM Catalog (611 results)
- Purpose: Bibliographic information for books, journals, and audiovisuals
- Example: Find textbooks and journals focused on hemoglobin disorders
- Activity: Students compile a reading list for a hypothetical course on globin biology
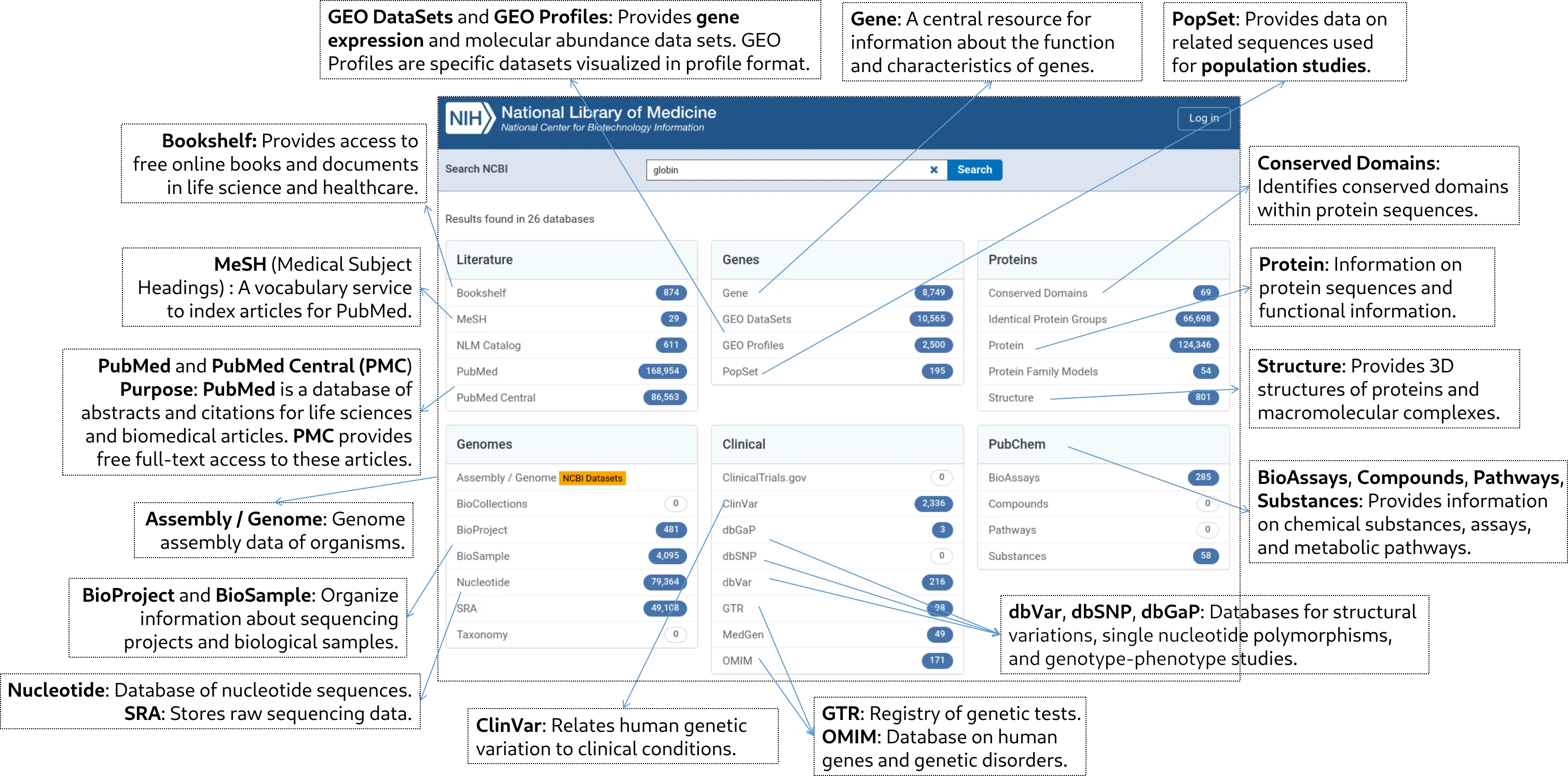
Section II. Genes Databases (25 minutes)
1. Gene (8,749 results)
- Purpose: Comprehensive information on genes from various species
- Example: Explore the human HBB (beta-globin) gene page
- Activity: Students compare globin genes across different species
2. GEO DataSets (10,565 results)
- Purpose: Repository for high-throughput gene expression data
- Example: Analyze a microarray dataset comparing normal and sickle cell erythrocytes
- Activity: Students interpret a simple gene expression heatmap related to globin expression
3. PopSet
- Purpose: Provides data on related sequences used for population studies.
- Use Case: Explore how globin gene sequences vary across populations.
- Exercise: Have students retrieve and compare globin gene variations in different populations using a PopSet entry.
Section III. Proteins Databases
1. Protein (124,346 results)
- Purpose: Comprehensive protein sequence and functional information
- Example: Analyze the sequence and structure of hemoglobin subunits
- Activity: Students use BLAST to compare globin protein sequences across species
2. Structure (801 results)
- Purpose: 3D macromolecular structure data
- Example: Visualize the 3D structure of oxyhemoglobin
- Activity: Students use a protein viewer to explore hemoglobin’s quaternary structure
3. Conserved Domains (69 results)
- Purpose: Database of protein domains and functional sites
- Example: Identify conserved domains in globin proteins
- Activity: Students compare domain structures of different globin family members
4. Protein Family Models (54 results)
- Purpose: Collection of protein family definitions
- Example: Examine the globin protein family model
- Activity: Students use protein family models to predict functions of hypothetical proteins
Section IV. Genomic Databases (25 minutes)
1. Assembly/Genome (NCBI Datasets)
- Purpose: Access to genome assemblies and annotations
- Example: Examine the genomic context of globin genes in the human genome
- Activity: Students use a genome browser to visualize the beta-globin locus
2. BioProject (481 results)
- Purpose: Central access point for project metadata
- Example: Examine a large-scale project on globin gene regulation
- Activity: Students summarize the objectives and data types of a globin-related BioProject
3. BioSample (4,095 results)
- Purpose: Biological source materials used in studies
- Example: Analyze sample information from a study on different hemoglobin variants
- Activity: Students design a BioSample submission for a hypothetical globin study
4. Nucleotide (79,364 results)
- Purpose: Collection of DNA and RNA sequences
- Example: Retrieve and compare globin gene sequences from different species
- Activity: Students perform a multiple sequence alignment of globin genes
5. SRA (49,108 results)
- Purpose: Raw sequencing data archive
- Example: Explore RNA-seq data from a study on globin gene expression during development
- Activity: Students analyze a small RNA-seq dataset to identify differentially expressed globin genes
6. BioCollections (0 results)
- Purpose: Information about biological collections and natural history museums
- Why no “globin” results: Focuses on organism-level collections, not molecular data
- Example: Explore a collection of marine organisms for potential novel globin research
- Activity: Students design a collecting expedition to study globin evolution in extreme environments
7. Taxonomy (0 results)
- Purpose: Hierarchical classification of organisms
- Why no “globin” results: Categorizes organisms, not genes or proteins
- Example: Trace the evolutionary history of organisms known to have unique globin variants
- Activity: Students create a phylogenetic tree of species with well-studied globin proteins
Section V. Clinical Databases
1. ClinVar (2,336 results)
- Purpose: Archive of relationships between human genetic variants and phenotypes
- Example: Investigate pathogenic variants in the HBB gene associated with beta-thalassemia
- Activity: Students research and present on a specific globin-related genetic disorder
2. OMIM (171 results)
- Purpose: Catalog of human genes and genetic disorders
- Example: Explore the OMIM entry for sickle cell anemia
- Activity: Students create a family pedigree for a globin-related genetic disorder
3. dbGaP (3 results)
- Purpose: Archive of genotype and phenotype interaction studies
- Example: Examine a genome-wide association study related to hemoglobin levels
- Activity: Students interpret basic GWAS results related to globin genes
4. GTR (98 results)
- Purpose: Genetic Testing Registry
- Example: Find genetic tests available for hemoglobinopathies
- Activity: Students create a patient information sheet for a specific globin-related genetic test
5. MedGen (49 results)
- Purpose: Organized information about human medical genetics
- Example: Explore the genetic basis of thalassemias
- Activity: Students create a concept map linking genetic variations to clinical presentations in globin disorders
6. ClinicalTrials.gov (0 results)
- Purpose: Registry of clinical studies
- Why no “globin” results: Likely due to specific search term limitations in the NCBI interface
- Example: Directly search ClinicalTrials.gov for “hemoglobin” or “sickle cell” studies
- Activity: Students design a hypothetical clinical trial for a new globin-related therapy
7. dbSNP (0 results)
- Purpose: Database of short genetic variations
- Why no “globin” results: Likely due to specific search term limitations in the NCBI interface
- Example: Search for SNPs in globin genes using gene names (e.g., HBB, HBA1)
- Activity: Students analyze the population frequency of a specific globin-related SNP
Section VI. Chemical and Assay Databases
1. BioAssays (285 results)
- Purpose: Archive of bioactivity screening data
- Example: Review assays testing compounds that affect hemoglobin oxygen affinity
- Activity: Students design a hypothetical bioassay for a globin-related research question
2. Substances (58 results)
- Purpose: Chemical substance information
- Example: Explore chemical data on heme and its derivatives
- Activity: Students create a concept map linking different substances involved in globin function
3. Compounds (0 results)
- Purpose: Information about chemical structures and their biological activities
- Why no “globin” results: Globin is a protein, not a small molecule compound
- Example: Search for compounds that interact with hemoglobin (e.g., 2,3-BPG)
- Activity: Students propose a novel compound that could potentially modify globin function
4. Pathways (0 results)
- Purpose: Biological pathway and interaction network information
- Why no “globin” results: “Globin” alone might not be recognized as a pathway term
- Example: Search for “heme biosynthesis” or “erythropoiesis” pathways
- Activity: Students create a simple pathway diagram showing globin synthesis and degradation
ClinVar vs OMIM: Key Differences
- Focus: CILVAR emphasizes clinical information relevant to rare diseases, while OMIM provides extensive genetic details about genes and phenotypes.
- Audience: CILVAR is geared more toward clinicians and healthcare providers needing practical information for patient care. In contrast, OMIM caters to researchers and geneticists focused on understanding genetic disorders at a molecular level.
Both databases are essential resources in their respective domains, contributing to the understanding and management of genetic disorders.
Accessing NCBI Data Using the rentrez Package in R
The National Center for Biotechnology Information (NCBI) provides vast amounts of biological data, including millions of scientific papers, genetic sequences, and species information. The Entrez system offers a powerful search engine and API for accessing this data. Rentrez is an R package that provides functions to interface with Entrez, simplifying data retrieval and analysis within your R sessions.
1. Installation
The rentrez package is available on CRAN (Comprehensive R Archive Network).
- To install it, type the following in your R console:
install.packages("rentrez")-
Once installed, load the package into your R session:
library(rentrez)
The rentrez package provides functions to explore available NCBI databases and search fields.
-
entrez_dbs(): Returns a list of all available NCBI databases.
-
entrez_db_summary(“database_name”): Provides a summary of a specific database, including its description and number of entries.
-
entrez_db_searchable(“database_name”): Returns a named list of available search fields for a specific database.
2. Identifying NCBI Databases
The entrez_dbs() function displays a list of all searchable databases in NCBI. You can use this list to determine which database to use for your queries.
entrez_dbs()Exploring Searchable Fields
The entrez_db_searchable() function reveals the searchable fields within a specific database. This is crucial for constructing accurate and targeted queries.
entrez_db_searchable(db = "snp") # Explore searchable fields in the SNP databaseAccessing Database Metadata with entrez_info()
The entrez_info() function retrieves metadata about a specific NCBI database.
entrez_info(db = "mesh")This output provides information like database name, description, fields, and links to other databases.
The entrez_db_summary**()** function retrieves a summary description about a specific NCBI database.
entrez_db_summary(‘pubmed’)
3. Searching NCBI Databases with entrez_search()
The entrez_search() function allows you to perform searches within a specific database. It takes two essential arguments:
-
db: The name of the database you want to search (e.g., “pubmed”, “protein”, “nuccore”).
-
term: The search term(s).
The function returns a list containing:
-
count: The total number of search results.
-
ids: A list of unique identifiers for matching records.
res <- entrez_search(db = "pubmed", term = "(PLoS Neglected Tropical Diseases[JOUR] AND 2015[PDAT])")res$count # Total resultsres$ids # IDs of matching records
By default, entrez_search() returns a maximum of 20 IDs. To retrieve more, use the retmax** argument**:
res <- entrez_search(db = "pubmed", term = "(PLoS Neglected Tropical Diseases[JOUR] AND 2015[PDAT])", retmax = 9999, use_history = TRUE)Example 2: To find papers in PubMed related to “COVID-19”, you would use:
covid_search <- entrez_search(db = "pubmed", term = "COVID-19")The covid_search object will contain a list of IDs (PMIDs in this case) that match your search criteria. You can access the IDs using:
covid_search$ids4. Exploring Search Results
To get more information about each record, use theentrez_summary() function:
covid_summs <- entrez_summary(db = "pubmed", id = covid_search$ids)This will return a list of summary records, each containing details about a specific PMID. You can use the extract_from_esummary function to extract specific fields from these records:
titles <- extract_from_esummary(covid_summs, "title")unname(titles)This will display the titles of the articles in the search results.
5. Downloading Data
The entrez_fetch() function is used to download data from NCBI databases. You need to provide the database (db), a list of IDs (id), and the desired data format (rettype).
For example, let’s do a search for “beta globin” term in the “Nucleotide (nuccore)” database:
globinprot <- entrez_search(db = “nuccore”, term = “beta globin”)
To fetch the sequences associated with a set of IDs in FASTA format:
seqs <- entrez_fetch(db = “nuccore”, id = globinprot$ids[1:3], rettype = “fasta”)
You can then save the retrieved data to a file:
write(seqs, "globin_sequences.fasta")Advanced: Using Web History for Large Queries
For large searches, you can use the use_history = TRUE option in entrez_search to store your search results on the NCBI server. This allows you to retrieve data in smaller batches, preventing API rate limits.
snp_search <- entrez_search(db = "snp", term = "Y[CHR] AND Homo[ORGN] NOT 10001:2781479[CPOS]", use_history = TRUE)
recs <- entrez_fetch(db = "snp", web_history = snp_search$web_history, retmax = 5, rettype = "xml", parsed = TRUE)Handling Large Queries with web_history and entrez_post()
NCBI limits the size of queries. To handle large datasets, use the web_history object, which stores lists of IDs on NCBI servers:
res <- entrez_search(db = "pubmed", term = "(PLoS Neglected Tropical Diseases[JOUR] AND 2015[PDAT])", retmax = 9999, use_history = TRUE)recs <- entrez_fetch(db = "pubmed", web_history = res$web_history, rettype = "xml", parsed = TRUE)Advanced: Linking Data across Databases with entrez_link()
One of the most powerful features of NCBI is the ability to link data across databases. The entrez_link() function allows you to retrieve linked records from different databases based on a specific record ID.
yfm <- entrez_search(db = "taxonomy", term = "yellow fever mosquito")yfm$ids # Get the ID of the "yellow fever mosquito"To retrieve sequences for this mosquito, you can link the taxonomy record to the genome database and then to the nuccore nucleotide database:
yfmlinks <- entrez_link(dbfrom = "taxonomy", id = yfm$ids, db = "genome")genlinkid <- yfmlinks$links$taxonomy_genome # Genome ID
yfmlinks2 <- entrez_link(dbfrom = "genome", id = genlinkid, db = "nuccore")nuclinkid <- yfmlinks2$links$genome_nuccore # Nucleotide IDThen, you can fetch the nucleotide sequences in FASTA format:
yfmfasta <- entrez_fetch(db = "nuccore", id = nuclinkid, rettype = "fasta")rentrez Exercises
These exercises will help you practice using the rentrez package in R.
Exercise 1: Finding PubMed Articles
-
Find all PubMed articles published in 2023 that mention “machine learning” and “cancer”.
-
Display the titles of the first 10 articles found.
-
Download the full text of the first article in XML format.
Exercise 2: Exploring GenBank
-
Find the GenBank entry for the human gene TP53.
-
Download the nucleotide sequence of the gene in FASTA format.
-
How many protein sequences are linked to this GenBank entry?
Exercise 3: Analyzing a PopSet Dataset
-
Search the PopSet database for datasets containing sequences of Drosophila melanogaster.
-
Find the dataset with the most sequences.
-
Download the FASTA sequences from that dataset.
-
(Optional) Use a package like ape to build a phylogenetic tree from the downloaded sequences.
Exercise 4: Comparing Database Information
-
Use entrez_db_searchable to list the available search fields for the “gene” and “pubmed” databases.
-
Are there any search fields that are common to both databases?
-
What are the specific search terms you can use in each database for the common fields?
Exercise 5: Using Web History
-
Perform a search in the “snp” database for all SNPs located on chromosome 1 in humans.
-
Use the web history object to download the first 20 SNPs in XML format.
-
Examine the XML structure of the downloaded data.
Rentrez Key Takeaways:
Getting Started with Rentrez
-
Exploring NCBI Databases:
-
Use entrez_dbs() to obtain a list of available NCBI databases.
-
Utilize functions like entrez_db_summary(), entrez_db_searchable(), and entrez_db_links() to learn more about each database.
-
Searching Databases: entrez_search()
-
Basic Searches:
-
Use entrez_search(db=“database_name”, term=“search_term”) to search a specific database.
-
The retmax argument controls the maximum number of returned IDs (defaults to 20).
-
-
Building Search Terms:
-
Use query[SEARCH FIELD] to target specific fields within a database.
-
Combine search terms using the Boolean operators AND, OR, and NOT.
-
Employ entrez_db_searchable() to identify searchable fields for a given database.
-
-
Using the “Filter” Field:
-
The “Filter” field allows you to refine searches based on specific criteria.
-
Explore available filtering terms using the “advanced search” tool on the NCBI website.
-
-
Precise Queries with MeSH Terms:
-
Medical Subject Headings (MeSH) provide a controlled vocabulary for highly specific searches.
-
Search for MeSH terms using entrez_search(db=“mesh”, term =…) to learn more about them.
-
Finding Cross-References: entrez_link()
-
Discovering Links:
-
Use entrez_link(dbfrom=“source_database”, id=“ID”, db=“target_database”) to find linked records in other databases.
-
Set db=“all” to retrieve links from all databases.
-
-
Narrowing Your Focus:
- Specify the db argument to target specific databases for linking.
-
External Links:
-
entrez_link(cmd=“llinks”) identifies external links, such as full-text article sources.
-
Use linkout_urls() to extract URLs from external links.
-
-
Multiple IDs:
-
Pass multiple IDs to entrez_link() to get links for all.
-
Use the by_id=TRUE argument to retain ID-specific links.
-
Getting Summary Data: entrez_summary()
-
Summary Records:
-
Use entrez_summary(db=“database_name”, id=“ID”) to retrieve a summary record for a given ID.
-
Explore elements within the returned object using the $ operator.
-
-
Multiple Records:
-
Pass multiple IDs to entrez_summary() to get summaries for each.
-
Utilize extract_from_esummary() to extract specific elements from multiple summary records.
-
Fetching Full Records: entrez_fetch()
-
Full Records:
- Use entrez_fetch(db=“database_name”, id=“ID”, rettype=“format”) to retrieve complete records in various formats (e.g., FASTA, XML).
-
FASTA Format:
- rettype=“fasta” retrieves sequences in FASTA format.
-
Parsed XML Documents:
-
rettype=“xml”, parsed=TRUE downloads and parses XML records.
-
Use XML::xmlToList(), XML::xpathSApply(), or XPath expressions to extract information from XML documents.
-
Using NCBI’s Web History Features
-
Posting IDs:
-
Use entrez_post(db=“database_name”, id=“ID”) to store IDs on the NCBI servers for later use.
-
The function returns a web_history object containing information for accessing the posted IDs.
-
-
Using Web History Objects:
-
Pass web_history objects to entrez_search(), entrez_summary(), and entrez_link() instead of IDs.
-
Use the use_history=TRUE argument in entrez_search() and entrez_link() to save search results as web_history objects.
-
Rate-Limiting and API Keys
-
Default Rate Limit:
- The NCBI limits users to 3 requests per second.
-
API Keys:
-
Register for a “my ncbi” account to obtain an API key for higher request limits.
-
Use the api_key argument in function calls or set the ENTREZ_KEY environment variable to take advantage of your API key.
-
-
Managing Rate Limits:
- Include Sys.sleep(0.1) before each request to the NCBI to prevent rate-limiting errors.
db vs dbform
db:
-
Indicates the target database: This argument tells rentrez which NCBI database you want to access.
-
Examples:
-
entrez_search(db=“pubmed”, term=“R Language”): Searches the PubMed database for articles about the R language.
-
entrez_link(dbfrom=“gene”, id=351, db=“nuccore”): Finds links to the nucleotide database (nuccore) for the gene with ID 351.
-
entrez_fetch(db=“nuccore”, id=linked_transripts, rettype=“fasta”): Fetches DNA sequences in FASTA format from the nucleotide database.
-
dbfrom:
-
Specifies the source database: This argument tells rentrez where the ID you’re providing comes from. This is essential for finding links between records from different databases.
-
Examples:
-
entrez_link(dbfrom=“gene”, id=351, db=“all”): Looks for links to all databases starting from the gene database (gene) using the gene ID 351.
-
entrez_link(dbfrom=“omim”, db=“clinvar”, cmd=“neighbor_history”, id=600807): Finds genetic variants in the ClinVar database related to asthma, using the OMIM ID 600807.
-
In Summary:
-
db tells rentrez where you want to go (target database).
-
dbfrom tells rentrez where you’re coming from (source database).
These arguments often work in tandem to connect records from different databases and explore complex relationships within NCBI data.