2. Downloading Data from Gene Expression Omnibus (GEO) using GEOquery
This tutorial will guide you through the process of downloading and accessing data from the Gene Expression Omnibus (GEO), a public repository of high-throughput data, using the GEOquery package in R.
1. What is GEO?
GEO is a valuable resource for researchers working with various types of biological data. It houses datasets from different studies, including:
-
Gene expression data: This is the most common type of data in GEO, representing gene activity levels in different cells or tissues.
-
Epigenetic data: This data relates to modifications to DNA or its associated proteins, influencing gene expression.
-
Other data types: GEO also stores data from various other high-throughput experiments, including microarrays, next-generation sequencing, and proteomics.
2. Understanding GEO Data Organization
Navigating GEO’s Structure
-
GEO Datasets: Datasets represent collections of biologically comparable GEO Samples. All samples within a dataset use the same platform.
-
GEO Profiles: Profiles provide the expression measurements for a single gene across all samples in a dataset.
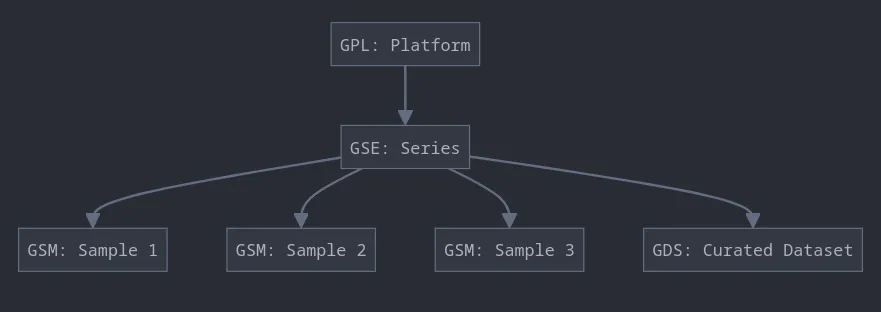
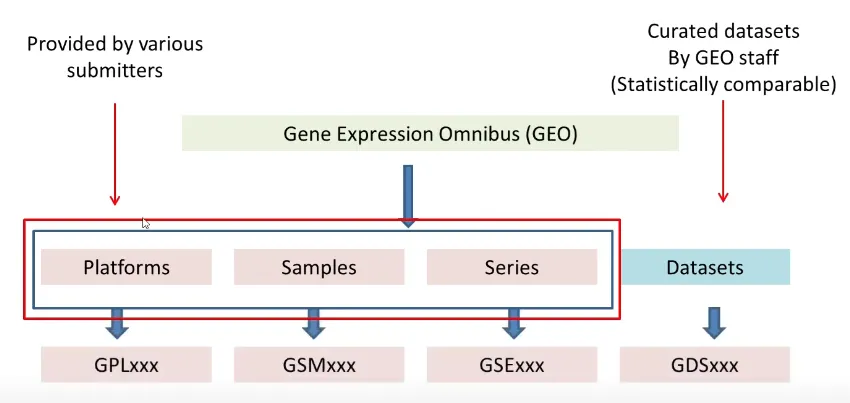
GEO is organized into four key entity types:
-
Platforms (GPLxxx): Describe the elements on an array (e.g., probes, antibodies) or the elements that can be measured in an experiment (e.g., SAGE tags, peptides).
-
Samples (GSMxxx): Represent individual experiments and contain information about the sample, its treatment, and the measurements obtained.
-
Series (GSExxx): Define sets of related samples, often representing a specific study or experiment.
-
Datasets (GDSxxx): Curated sets of GEO samples that are biologically and statistically comparable, often used for analysis and visualization.
- Dataset Example: GSE123456 (Human pancreatic cancer gene expression)
- Dataset Example: GSE987654 (Type 2 diabetes gene expression, healthy vs. patients)
- Dataset Example: GSE789012 (Human blood gene expression, healthy vs. COVID-19 patients)
3. Getting Started with the GEOquery Package
The GEOquery package offers a comprehensive set of functions to help you process, analyze, and visualize data from GEO. Explore the documentation for more advanced techniques.
Install GEOquery: If you haven’t already, install the GEOquery package in R.
if (!require("BiocManager", quietly = TRUE)) install.packages("BiocManager")
BiocManager::install("GEOquery")Load the package:
library(GEOquery)4. Accessing GEO Data
The main function for retrieving data from GEO is getGEO(). You can use it with the following syntax:
# Get a GDS datasetgds <- getGEO("GDS507")
# Get a GSM samplegsm <- getGEO("GSM11805")
# Get a GSE seriesgse <- getGEO("GSE781", GSEMatrix = FALSE)Note that you typically provide the GEO accession number to getGEO(), which is a unique identifier for each entry in GEO. If you don’t have an internet connection, you can use the filename argument to access data from a local file.
2.1 Downloading a GEO Series (GSE)
To download a series, such as GSE21653 (be aware this is a 20 Mb download!), use the following command:
gse <- getGEO("GSE21653", GSEMatrix = TRUE)show(gse)This downloads the entire series, including the expression matrix. The show() function provides an overview of the downloaded series.
Explore the Data:
GEOquery provides specific data structures for each of the four GEO entity types:
-
GDS, GSM, and GPL classes: These classes share a similar structure, containing a metadata header (derived from the SOFT format) and a GEODataTable. The GEODataTable stores the actual measurements or values, along with column descriptions. You can access the metadata using Meta(object) and the data table using Table(object).
-
GSE class: The GSE class represents a series and contains lists of GSM and GPL objects within it. You can access the lists using GSMList(object) and GPLList(object).
Example:
# Access metadata for a GSM objecthead(Meta(gsm))
# Access data table for a GSM objectTable(gsm)[1:5,]
# Access column descriptionsColumns(gsm)
# Access GSM list for a GSE objectnames(GSMList(gse))4. Working with Raw Data (Supplementary Files)
Sometimes, you might need to download raw data files associated with a study. This is how you can do it:
-
Get Supplementary Files:
supplementaryFiles <- getGEOSuppFiles("GSE12345") -
Access the Downloaded Files:
The getGEOSuppFiles function will download the files into a TAR archive. You can then extract the files from the archive.
-
Accessing Raw Data from GEO
The getGEOSuppFiles() function allows you to download the raw data associated with a specific GEO accession (e.g., .CEL files, images). By default, it creates a directory to store the downloaded files.
# Download raw data for a GSM accessiongetGEOSuppFiles("GSM11805")
Example: Downloading Raw Data from a Microarray Study
-
Suppose you want to download the raw data (CEL files) from a study using Affymetrix microarrays with the GEO series accession number GSE12345.
supplementaryFiles <- getGEOSuppFiles("GSE12345") -
**You can then use untar to extract the files from the TAR archive. This will create a directory containing the CEL files.
untar("GSE12345_RAW.tar", exdir = "GSE12345_RAW")
5. Converting GEO Data to Bioconductor Objects
5.1 Converting GDS to ExpressionSet:
The GDS2eSet() function converts a GDS object to a Bioconductor ExpressionSet object, which is commonly used for microarray analysis.
# Convert GDS to ExpressionSeteset <- GDS2eSet(gds, do.log2 = TRUE)
# Explore the ExpressionSeteset5.2 Converting GDS to MAList:
The GDS2MA() function converts a GDS object to a MAList object from the limma package. MAList is another common data structure for microarray analysis, offering the ability to store gene annotation information.
# Get platform annotationgpl <- getGEO("GPL97")
# Convert GDS to MAListMA <- GDS2MA(gds, GPL = gpl)
# Check the class of the MAList objectclass(MA)5.3 Converting GSE to ExpressionSet:
Converting a GSE object to an ExpressionSet can be more complex, as it may contain multiple samples from different platforms. The process typically involves filtering the GSMList to include only samples from the desired platform and constructing the data matrix manually.
# Filter GSMList for a specific platformgsmlist <- Filter(function(gsm) {Meta(gsm)$platform_id == 'GPL96'}, GSMList(gse))
# Extract data values from filtered GSMListdata.matrix <- do.call('cbind', lapply(gsmlist, function(x) { tab <- Table(x) mymatch <- match(probesets, tab$ID_REF) return(tab$VALUE[mymatch])}))
# Create ExpressionSet object from data matrixeset2 <- new('ExpressionSet', exprs = data.matrix, phenoData = pheno)
# Examine the ExpressionSeteset26. Accessing Phenotypic Information
6.1 Using the getGEO() Function
The getGEO() function allows you to access phenotypic information stored within a GSE object.
dim(pData(gse[[1]])) # Shows the dimensions of the phenoData objecthead(pData(gse[[1]])[, 1:3]) # Displays the first few rows and columns of the phenoData objectKeep in mind this method downloads the entire GSE matrix, which can be resource-intensive.
6.2 Using the getGSEDataTables() Function
Some GSEs come with separate data tables containing sample information. You can access these tables using the getGSEDataTables() function.
df1 <- getGSEDataTables("GSE3494")lapply(df1, head)The lapply() function iterates through the data tables and displays the first few rows of each.